We recently released valid_data, a gem to check whether the rows in your database are valid according to your ActiveRecord validations. It worked by sequentially scanning your tables in order to check if the model instances were still valid over time. While this is pretty straightforward, we're essentially being naive in the sense that we could actually process some tables in parallel. In this post we'll be discussing the concept and implementation of parallelization that we chose to implement in the valid_data gem. We'll also show how it dramatically improves performance in certain cases over the sequential library.
What are we trying to solve?
The problem that we're trying to solve has some characteristics that we can take advantage of. Taking a closer look at the data, we can break the problem into smaller parts (tables), process them independently and then combine the solutions to show the final result.
Below we distribute models M1, M2, M3, M4 and M5 between 3
buckets. P1, P2 and P3 will then process the models
sequentially. First, P1 will sequentially process M1 and M4,
P2 will do the same for M2 and M5, and so on and so forth.
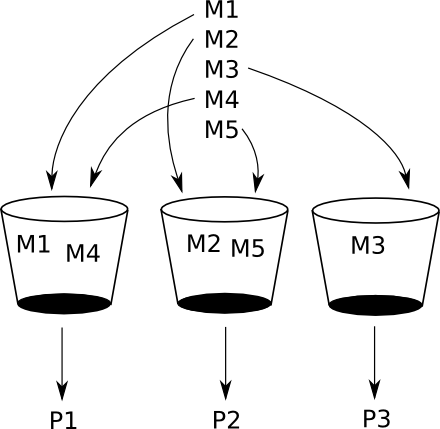
We could even take a step further and have 5 buckets each containing just one model if we know we have enough capacity for that. We'll take a look at that later in the benchmarks section.
Implementation
In Unix-like operating systems there is a concept called fork where a process can create a copy of itself. This is a very cheap and fast process that can be used to implement parallelization. After forking a process, we have the original parent process and the new copy commonly called child process. We're not limited to create only one child, but as many as we want. In general, most libraries allow users to choose how many children will be forked making easier to customize according to the number of CPUs in their machine. We implemented that by accepting the number of children as an argument for our task:
rake validate_records[30,8]
In the execution above, we ask it to fork 8 child processes so that the processing of models will be distributed between them. This is more or less translated to the following code:
@readers = fork(num_of_processes.to_i, &worker)
The @readers variable will hold the communication mechanism between
child processes and their parent process. The mechanism by which
children can communicate with their parents is known as a "pipe."
reader, writer = IO.pipe
In order to keep the parent process aware of any message sent by their children we create a thread that will periodically check if there is any message from any child's pipe. Here is a simplified version:
Thread.new do
loop do
io = IO.select(readers.values)
((io && io.first) || []).each do |reader|
yield reader
end
end
end
Finally we tell the parent process to wait for its children to finish their processing with:
wait(pids(@readers))
There's a gotcha here. We have to wait as many times as we forked. In
other words, you have to call Process.wait as many times as you call
Process.fork. That's why we extract the pids (Process IDs) from the
readers in order to wait for all children processes. You can find the
complete code
here.
Benchmarks
After implementing parallelization with the concept previously discussed, it's time to measure if it is indeed worth going in this direction. We've designed our schema with the following tables:
Model | Total
------------------------------
SubPage | 22000
StaticPage | 22000
Essay | 40000
News | 40000
Page | 40000
Article | 496687
Post | 572000
Each one one has two columns: title and body. You can find the
entire schema, benchmark script and machine details
here. With the
schema above we've run some benchmarks to compare:
- Sequential execution
- Parallel execution with 2 children
- Parallel execution with 5 children
- Parallel execution with 7 children
After 5 iterations we were able to see how the time is decreasing as we increase the number of processes:
user system total real
Sequential 555.080000 5.120000 560.200000 (606.937533)
Parallel 2x 0.830000 0.150000 359.200000 (236.735275)
Parallel 5x 0.820000 0.290000 317.530000 (185.183759)
Parallel 7x 0.760000 0.290000 320.680000 (175.603909)
In a benchmark with 10 iterations we see a similar speedup:
user system total real
Sequential 1136.120000 9.940000 1146.060000 (1240.820179)
Parallel 2x 1.600000 0.340000 643.410000 (425.282860)
Parallel 5x 1.630000 0.470000 639.670000 (362.135289)
Parallel 7x 1.580000 0.540000 678.880000 (358.012563)
We can see the processing time has dropped around 70% when comparing the sequential execution with parallel execution using 7 child processes. That's really impressive and the reason behind it is pretty straightforward. Before processing the tables we sort them by size so that the bigger tables will be at the end of the list. We take advantage of this and distribute big tables between different processes which in turn allows them to be processed in parallel:
$ ps aux | grep [V]alid
jivago 25812 [ValidData-0.0.1] Manager [25838, 25841, 25844, 25847, 25851, 25855, 25859]
jivago 25838 [ValidData-0.0.1] Processing ["SubPage"]
jivago 25841 [ValidData-0.0.1] Processing ["StaticPage"]
jivago 25844 [ValidData-0.0.1] Processing ["Essay"]
jivago 25847 [ValidData-0.0.1] Processing ["News"]
jivago 25851 [ValidData-0.0.1] Processing ["Page"]
jivago 25855 [ValidData-0.0.1] Processing ["Article"]
jivago 25859 [ValidData-0.0.1] Processing ["Post"]
We can see above the models' distribution among processes. In this scenario, each model is being examined by a different process. After some time the smaller tables will be finished and we can see the bigger ones still being processed in parallel:
$ ps aux | grep [V]alid
jivago 25812 [ValidData-0.0.1] Manager [25855, 25859]
jivago 25855 [ValidData-0.0.1] Processing ["Article"]
jivago 25859 [ValidData-0.0.1] Processing ["Post"]
Conclusion
We've seen that breaking the problem into smaller parts allowed us to process them independently with parallel execution. Parallelization has dropped the time by 70%. While this schema is not a real production system, chances are you'll probably have much more bigger tables and that's exactly where this approach can be of the most benefit.
As future work it may be worth distributing a gigantic table between processes which in practice would be translated in more than one process checking different parts of the same table. Other scenario worth investigating would be checking how the parallel processing is affected when there are associations between models. In this case, while processing a table we will end up retrieving data from other tables and that may affect the overall speed since now the parts are not truly independent from each other. For scenarios like that figuring out a good distribution of group of tables may help to surpass the dependency problem.
Please let us know if you have any question or ideas to improve the solution or the code. We hope this new approach will bring velocity to you in your daily job!






